Discovering targets and biomarkers for immunotherapies
Immunological treatments for cancer encompass a broad category of established and emerging therapies. While immunotherapies are known to eradicate even advanced tumors in which other treatments have failed, many patients experience little to no benefit. A better understanding of tumor immunity, better target antigens and better markers for response and resistance are needed. Tumor DNA- and RNA-sequencing play an important role in this research. Here we discuss applicable computational analyses of NGS data with emphasis on T-cell-related therapies.
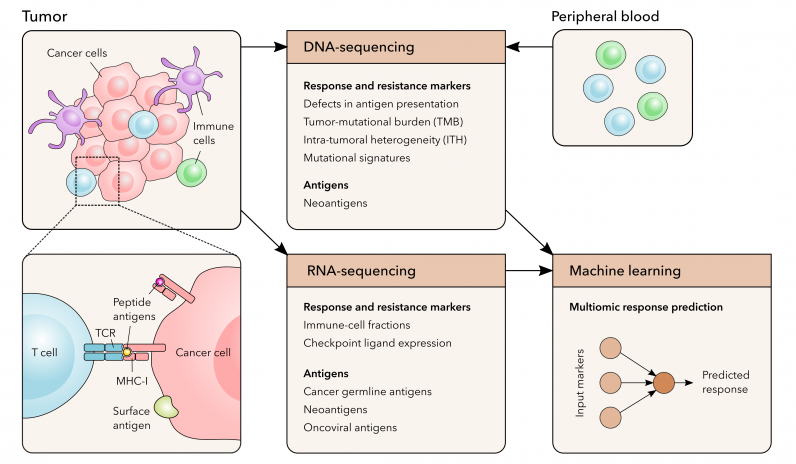
Tumor immunity and its evasion
Our immune system is, in principle, capable of killing cancer cells. By targeting tumor-associated and tumor-specific antigens (TAAs and TSAs), lymphocyte receptors – T-cell receptors and antibodies – enable the immune system to detect cancer cells and eliminate them.
However, the immune system often fails to eliminate cancer, for reasons such as:
- Poor immune-cell infiltrability of tumor due to physical, chemical and biological barriers
- A lack or scarcity of cancer antigens
- Defects in antigen presentation
- A lack of naturally developed lymphocyte receptors against cancer antigens
- The expression of immune-inhibitory signaling molecules by the tumor or other cells in the tumor microenvironment (TME).
Under the pressure imposed by both normal immune surveillance and immunotherapy, tumors regularly evolve to exploit these immuno-evasive mechanisms. Tumor evolution, thus, arguably poses the greatest challenge in developing effective immunotherapies.
While tumor types generally vary in terms of their immunogenicity, it can be helpful to look beyond the tumor's type or tissue of origin and focus on its immune microenvironment. This approach aids in developing treatments across different cancer types, as long as crucial immunological properties are shared.
Tumors can be classified by their immune microenvironment into hot and cold tumors (exhibiting high and low immune infiltration, respectively) and into more granular immune archetypes that take into account the types and activities of immune cells.
Types of immunotherapy
Immunological cancer therapies are designed to enhance the immune system's ability to target and destroy cancer cells. These therapies encompass various approaches, including the use of small molecules, antibodies, living immune cells, vaccines, and oncolytic viruses. The primary mechanism often relies on cytotoxic T cells attacking and eliminating cancer cells. Immunotherapies include, but are not limited to:
- Immune checkpoint inhibition (ICI): ICI therapies aim to reactivate lymphocytes, particularly cytotoxic T cells, whose activity has been suppressed by checkpoint receptors. Checkpoint receptors like PD1 and CTLA4 normally exist to prevent excessive immune responses, including autoimmunity. However, cancer cells can exploit these mechanisms to evade the immune system's detection. Most checkpoint inhibitors are monoclonal antibodies targeting either the receptors themselves or their ligands.
- Tumor-infiltrating lymphocyte (TIL) treatment: TIL treatment involves the extraction of lymphocytes, T cells in particular, from within the tumor. These cells are then activated and expanded outside the body before being reintroduced into the patient. The rationale behind this approach is that T cells already present within the tumor are more likely to recognize tumor-specific antigens, and increasing their numbers enhances their anti-cancer activity.
- TCR T-cell therapy: Like TIL treatment, TCR (T-cell receptor) T-cell therapy involves ex vivo T-cell expansion. However, it does not rely on the natural development of immune receptors against tumor antigens. Instead, it involves the genetic engineering of a patient's own T cells to express receptors specifically designed to target known tumor antigens. These antigens might include cancer germline antigens (CTAs) or antigens associated with certain viral infections (onco-viral antigens).
- CAR T-cell therapy: The third type of adoptive cell therapy on this list, CAR T-cell therapy shares similarities with TCR T-cell therapy but employs a different engineered receptor known as chimeric antigen receptor (CAR). CARs differ from TCRs in that they recognize individual proteins on the cell surface rather than MHC (major histocompatibility complex)-epitope complexes, which TCRs are restricted to.
- Bispecific antibodies: An antibody engineered to target a cancer antigen through one of its arms and a T-cell marker through the other one can be used to bring T-cells into contact with tumor cells, facilitating their recognition and killing.
- Therapeutic cancer vaccines: Therapeutic cancer vaccines, in contrast to traditional preventive vaccines used against cancer-causing viruses, aim to train the immune system to recognize antigens present in existing tumors. These vaccines may involve delivering tumor cell lysates, antigen-encoding DNA or mRNA, antigen proteins, parts of antigens (peptides), or dendritic cells carrying the antigen into the patient. Eventually, antigen-presenting dendritic cells license T-cells with suitable TCRs to engage in anti-cancer activity.
- Oncolytic viruses: Oncolytic viruses are engineered to preferentially infect cancer cells and engage in cytolytic activity within them. While oncolytic viruses must avoid immune destruction before locating and infecting cancer cells, the immune system also plays a part in their mechanism of action: both the viral infection itself and resulting cancer cell death enable immune cells to better recognize and kill cancer cells on their own. Thus, oncolytic viruses may effectively function as therapeutic vaccines.
Discovering cancer antigens
The development of many immunotherapies relies on the discovery of suitable cancer antigens. These therapies cover CAR-T and TCR T-cell therapies, as well as cancer vaccines and bispecific T-cell engagers.
Types of cancer antigens
Cancer antigens can be categorized into several types:
- Neoantigens: These are proteins affected by somatic mutations. Neoantigens are often unique to each patient and are particularly relevant for patient-specific therapies.
- Cancer germline antigens (CGAs): Also known as cancer testis antigens, CGAs are endogenous proteins normally restricted to germline cells. They are frequently upregulated in cancers and are important antigens for immunotherapies due to their association with cancer, immunogenicity, and independence from mutations.
- Oncoviral antigens: These are proteins expressed in virally-induced cancers. They are specific to cancer and are highly immunogenic, making them desirable targets for immunotherapies. However, they are limited to the minority of cancers associated with viruses.
- Other antigens: These include products of aberrant gene expression in cancer, such as aberrant splice variants, human endogenous retroviral (HERV) protein expression (epigenetically suppressed in normal tissue), and long non-coding RNAs (lncRNAs) aberrantly translated in cancer.
Antigen discovery with RNA-sequencing
RNA-sequencing data, obtained from an individual patient's tumor biopsy or large cancer transcriptome databases, can be used to identify candidate antigens. When evaluating these candidates, several desirable properties should be considered:
- Cancer specificity: Antigen expression should be specific to cancer cells to minimize the potential for therapy-induced toxicity.
- Recurrence of expression: Identifying antigens with recurrent expression across different tumors maximizes the therapy's applicability.
- Oncogenicity: This property can be indirectly assessed using computational approaches or through separate experiments designed to evaluate it. An oncogenic antigen is desirable as it reduces the tumor's evolutionary pressure to downregulate its expression during therapy.
- Immunogenicity: A required property particularly important for neoantigens: computational tools exist to predict specific neoepitopes resulting from a mutation and to assess their immunogenicity. Immunogenic peptides can also be identified indirectly by studying the TCRs of expanded T-cell clones in cancer patients, using TCR sequencing in a bulk or single-cell setting.
- Surface expression: For treatments relying on surface antigens, such as CAR T therapy, it's essential to ensure that the gene product is expressed on the cell surface. This can be determined through computational predictions or annotations based on surface protein databases.
It should be noted that transcriptome-based analyses provide direct evidence only of transcription – not of translation, antigen presentation or surface expression. Proteomic and immunopeptidomic mass-spectrometry analyses can be used to gather further evidence of the latter.
Discovering biomarkers for immunotherapy
The most important challenges in the development and application of immunotherapies are related to identifying patients who are most likely to benefit from treatment.
Discovering novel biomarkers for treatment response requires large sample sizes, and many known markers have thus been discovered – and may be specific to – the most widely applied immunotherapies, namely ICIs. While many biomarkers may be applied across different types of immunotherapies, some are necessarily treatment-specific, such as the expression of a specific target antigen.
Known and suggested molecular biomarkers include:
- Expression of target antigens in the tumor: This applies to antigen-specific treatments and can be assessed through immuno-assays against the antigen or on the transcript level from RNA-seq data.
- Tumor mutational burden (TMB): The overall mutational burden reflects the amount of neoantigens, and thus the immunogenicity of a tumor. TMB is estimated from tumor DNA sequencing data, either whole-genome or, more commonly, exome sequencing (WES) data.
- Neoantigen load: The neoantigen load is a more refined metric of immunogenicity compared to TMB. Computational predictors are used to estimate the amount of neoepitopes caused by mutations, taking into account the patient's HLA (human leukocyte antigen) alleles, which affect the types of epitopes presented.
- Mutational signatures: The spectrum of mutation types (e.g., different base substitutions, considered together with the mutation-flanking bases) in a tumor genome inform about the mutations’ source. Mutation spectra, or signatures, associated with processes such as mismatch repair deficiency (MMR) and APOBEC-induced mutagenesis, have been associated with immunotherapy response.
- Intratumoral heterogeneity (ITH): ITH measures mutational diversity among cancer cells. Highly heterogeneous tumors pose challenges for immunotherapies, as resistant subclones may expand rapidly, even if they are rare before treatment.
- Tumor-infiltrating immune cells: Tumor RNA-sequencing data, including single-cell and spatial transcriptomics, can be used to estimate the types and fractions of immune cells in the tumor microenvironment. Specific cell types are associated with immunogenic and immune-suppressive tumors.
- Expression of checkpoint ligands: Similar to antigen expression, immune checkpoint ligands (and receptors) can be quantified through immuno-assays or RNA-sequencing.
- HLA alleles and their diversity: The alleles of HLA genes, which are extremely polymorphic, can be detected from DNA-sequencing data using specialized tools. Each of the three canonical HLA-I genes (HLA-A, B and C) encodes for a HLA-I complex (also called MHC-I) together with a gene called beta-2 microglobulin (B2M). The diversity of HLA-I alleles affects the diversity of epitopes a cell presents, and a measure of this allelic diversity (HLA evolutionary divergence; HED) can thus be used as a biomarker for immunogenicity.
- Defects in antigen presentation: Loss of antigen presentation is a common resistance mechanism to cellular, or T-cell-mediated immunity. Complete loss of HLA-I expression may be detected by innate lymphocyte receptors and trigger TCR-independent killing of cancer cells by T and natural killer (NK) cells. For this reason, reduction in the number or diversity of presented antigens, rather than complete loss of antigen presentation, is commonly seen. Loss-of-function mutations, including genomic deletions and loss-of heterozygosity (LOH), may affect HLA-I alleles, B2M and a number of other genes involved in promoting and enabling HLA-I expression and antigen presentation. Somatic mutation analysis with HLA genes is particularly demanding due to their polymorphisms.
Many of these DNA- and RNA-sequencing-based markers of treatment response and resistance measure slightly different facets of tumor immunogenicity. They are likely to correlate with each other, and when considered together, the predictive power of certain markers may render other markers uninformative.
For these reasons, identifying the best combination of markers for an integrative predictive model may be more useful than attempting to identify the best single marker. With large datasets, machine learning models enable just that: the discovery of more complex and more predictive signatures. Importantly, accounting for multiple markers simultaneously may also increase the model's applicability to a wider range of tumor types.
Summary
DNA-sequencing enables identifying mutation-related measures of tumor immunogenicity and genomic defects in genes encoding the antigen presentation machinery.
RNA-sequencing applies particularly to identifying expressed antigens and immune infiltrates.
Machine learning allows combining different types of markers to discover complex response-predicting signatures.

Learn more
Our services and expertise
- Bioinformatics for cancer research
- Bioinformatics for immunology
- Bioinformatics for drug developmentTumor clonality analysisPredicting treatment response with machine learning
References and case studies
- Characterizing a Cancer Immunotherapy Target with Genomic Data (CDR-Life)Cancer risk prediction with machine learning (Karolinska Institute)
- Multiplex immunofluorescence analysis (Uppsala University)
Selected publications from our customers
- Chang, Y. T. et al. (2024). MHC-I upregulation safeguards neoplastic T cells in the skin against NK cell-mediated eradication in mycosis fungoides. Nature communications, 15(1), 752. https://doi.org/10.1038/s41467-024-45083-8
- Adebamowo, S. N. et al. (2024). Genome, HLA and polygenic risk score analyses for prevalent and persistent cervical human papillomavirus (HPV) infections. European journal of human genetics : EJHG, 10.1038/s41431-023-01521-7. Advance online publication. https://doi.org/10.1038/s41431-023-01521-7
- Karihtala, P. et al. (2023). Mutational signatures and their association with survival and gene expression in urological carcinomas. Neoplasia (New York, N.Y.), 44, 100933. Advance online publication. https://doi.org/10.1016/j.neo.2023.100933
- Backman, M. et al. (2023). Spatial immunophenotyping of the tumour microenvironment in non-small cell lung cancer. European journal of cancer (Oxford, England : 1990), 185, 40–52. https://doi.org/10.1016/j.ejca.2023.02.012
- Mezheyeuski, A. et al. (2023). An immune score reflecting pro- and anti-tumoural balance of tumour microenvironment has major prognostic impact and predicts immunotherapy response in solid cancers. EBioMedicine, 88, 104452. Advance online publication. https://doi.org/10.1016/j.ebiom.2023.104452
- Tusup, M. et al. (2022). Epitranscriptomics modifier pentostatin indirectly triggers Toll-like receptor 3 and can enhance immune infiltration in tumors. Molecular therapy : the journal of the American Society of Gene Therapy, 30(3), 1163–1170. https://doi.org/10.1016/j.ymthe.2021.09.022
- Ribeiro, R. et al. (2022). Synchronous Epidermodysplasia Verruciformis and Intraepithelial Lesion of the Vulva is Caused by Coinfection with α-HPV and β-HPV Genotypes and Facilitated by Mutations in Cell-Mediated Immunity Genes. Preprint at https://doi.org/10.21203/rs.3.rs-1991512/v1
- Madonna, G. et al. (2021). Clinical Categorization Algorithm (CLICAL) and Machine Learning Approach (SRF-CLICAL) to Predict Clinical Benefit to Immunotherapy in Metastatic Melanoma Patients: Real-World Evidence from the Istituto Nazionale Tumori IRCCS Fondazione Pascale, Napoli, Italy. Cancers, 13(16), 4164. https://doi.org/10.3390/cancers13164164
- Simao, F. et al. (2021). SRF-CLICAL: an approach for patient risk stratification using random forest models. bioRxiv 2021.06.22.448514; doi: https://doi.org/10.1101/2021.06.22.448514
- Gurvich, O. L. et al. (2020). Transcriptomics uncovers substantial variability associated with alterations in manufacturing processes of macrophage cell therapy products. Scientific reports, 10(1), 14049. https://doi.org/10.1038/s41598-020-70967-2
Selected publications from our team
- Fotakis, G. et al. (2024). Conventional therapy induces tumor immunoediting and modulates the immune contexture in colorectal cancer. bioRxiv 2024.08.21.608938; doi: https://doi.org/10.1101/2024.08.21.608938
- Caronni, N. et al. (2023). IL-1β+ macrophages fuel pathogenic inflammation in pancreatic cancer. Nature, 623(7986), 415–422. https://doi.org/10.1038/s41586-023-06685-2
- Mehtonen, J. et al. (2020). Single cell characterization of B-lymphoid differentiation and leukemic cell states during chemotherapy in ETV6-RUNX1-positive pediatric leukemia identifies drug-targetable transcription factor activities. Genome medicine, 12(1), 99. https://doi.org/10.1186/s13073-020-00799-2
- Dufva, O. et al. (2020). Immunogenomic Landscape of Hematological Malignancies. Cancer cell, 38(3), 380–399.e13. https://doi.org/10.1016/j.ccell.2020.06.002
- Lehtipuro, S. et al. (2019). Modes of immunosuppression in glioblastoma microenvironment. Oncotarget, 10(9), 920–921. https://doi.org/10.18632/oncotarget.26643
- Escobar, G. et al. (2018). Interferon gene therapy reprograms the leukemia microenvironment inducing protective immunity to multiple tumor antigens. Nature communications, 9(1), 2896. https://doi.org/10.1038/s41467-018-05315-0
Norelli, M. et al. (2018). Monocyte-derived IL-1 and IL-6 are differentially required for cytokine-release syndrome and neurotoxicity due to CAR T cells. Nature medicine, 24(6), 739–748. https://doi.org/10.1038/s41591-018-0036-4
- Luoto, S. et al. (2018). Computational Characterization of Suppressive Immune Microenvironments in Glioblastoma. Cancer research, 78(19), 5574–5585. https://doi.org/10.1158/0008-5472.CAN-17-3714
- Havunen, R. et al. (2018). Abscopal Effect in Non-injected Tumors Achieved with Cytokine-Armed Oncolytic Adenovirus. Molecular therapy oncolytics, 11, 109–121. https://doi.org/10.1016/j.omto.2018.10.005
Contact us
Leave us a message if you are interested in contracting us to get your data analyzed.
