Unraveling the pathogenesis of bovine mastitis with bacterial WGS analysis
A research group at the University of Helsinki, Faculty of Veterinary Medicine, is studying the pathogenesis of bovine mastitis. For their recently published study, Genevia Technologies analyzed bacterial whole-genome sequencing data to help understand whether putative virulence genes in Staphylococcus aureus impact the clinical outcome of bovine mastitis.
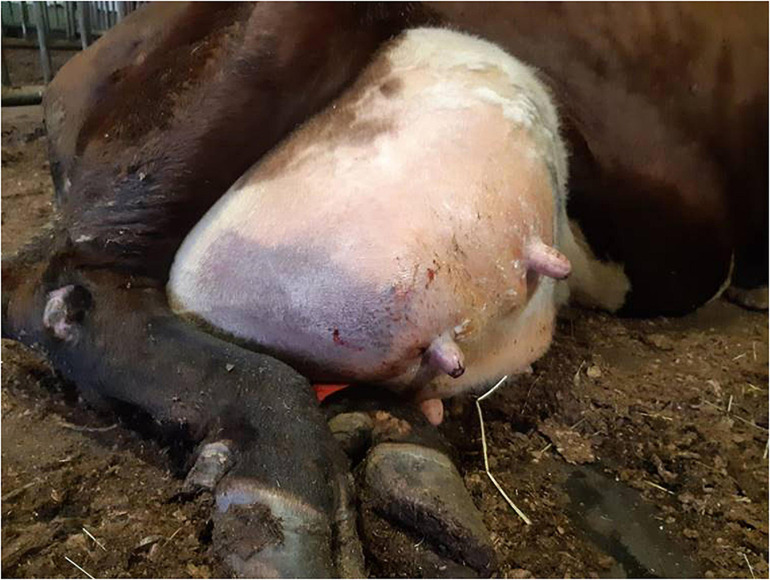
A dairy cow suffering from a peracute gangrenous S. aureus mastitis, photo by Mari Niemi
Bovine mastitis is the most common disease in dairy herds, and roughly one in every five Finnish dairy cows receive antimicrobial drugs for mastitis. The severity of the disease varies from mild, subclinical mastitis to severe, peracute gangrenous mastitis that can cause necrosis of the affected udder quarter and even lead to the death of the cow.
The mastitis research group at the University of Helsinki consists of microbiologist PhD Silja Åvall-Jääskeläinen; docent, DVM Suvi Taponen; DVM Joanna Koort and Associate professor, DVM Heli Simojoki. Their previous study searched for correlations between bovine mastitis types and virulence gene profiles in different Staphylococcus species. In this follow-up study, they focused on Staphylococcus aureus, intending to determine whether the presence or absence of genes encoding the putative virulence factors was associated with the clinical outcome of bovine S. aureus mastitis.
– The debate has been whether the severity of the Staphylococcus aureus mastitis depends on pathogen-related factors such as virulence genes or on the host’s defense mechanisms. We hypothesized that by comparing Staphylococcus aureus isolates gathered from subclinical and severe peracute mastitis, we could detect if the S. aureus strains impact the clinical outcome. We had collected bacterial samples over the years and sequenced the bacterial genomes, and now needed to find someone to analyze the WGS data. Genevia Technologies was chosen as our bioinformatics partner after our sequencing partner recommended them, says Dr. Taponen.
In search of an experienced and hassle-free bioinformatics provider
The research group says that the main criterion for selecting the bioinformatics partner was the overall smoothness of the bioinformatics workflow. Dr. Koort elaborates:
– The smoothness and easiness we were looking for were supported by our other criteria: the perfect partner for us would have the professionalism and substantial expertise to handle the bioinformatics workflow independently.
– And the ability to communicate in Finnish was considered a big plus, Dr. Åvall-Jääskeläinen continues.
The researchers agree in unison that hiring Genevia Technologies for the job gave them good value for money and that they are now one step closer to unraveling the pathogenesis of bovine mastitis.
– The careful data analysis showed us that there is no statistically significant correlation between the mere presence of any virulence genes and the severity of the disease. Therefore, future studies should focus on the allelic variation and especially different regulation and thus expression in the virulence genes together with the host’s genetics and immune defense, summarizes Dr. Suvi Taponen.
A smooth ride with Genevia Technologies from planning to publishing
Just one teleconference was all it took to agree on the data analysis workflow, which involved assembling and annotating genomes as well as pangenome, phylogenetic and gene ontology analyses. After the first meeting, the collaboration proceeded just as smoothly as the group had hoped for – or even more so. None of the researchers could think of any areas for improvement in Genevia’s bioinformatics service.
– In our experience, outsourcing was efficient, and the results were provided fast. Since everything went without a hassle with Genevia Technologies, we were able to focus on our job and didn’t need to worry about the computational experiments, Dr. Koort says.
Dr. Silja Åvall-Jääskeläinen wants to highlight the importance of a well-written project report:
– Genevia’s bioinformatician, Dr. Tommi Rantapero, had written such a comprehensive project report that I, a non-bioinformatician, could write the Materials and Methods section myself based on his report, which contained all the relevant citations. When submitting the paper, the bioinformatics section remained untouched.
– The deadlines were met beautifully, the quality of the work was spotless, and we had zero hiccups with our communication. So what is there not to recommend? We would choose Genevia Technologies again in a heartbeat, says Dr. Åvall-Jääskeläinen as the other members of the research group nod in agreement.
Published research from this collaboration
- Åvall-Jääskeläinen, S. et al. (2021). Genomic Analysis of Staphylococcus aureus Isolates Associated With Peracute Non-gangrenous or Gangrenous Mastitis and Comparison With Other Mastitis-Associated Staphylococcus aureus Isolates. Frontiers in microbiology, 12, 688819. https://doi.org/10.3389/fmicb.2021.688819
Contact us
Leave us a message below if you would like to hear how we can help you.
